A Spin-Glass Metabolic Hamiltonian optimized by Quantum Annealing Reveals Thermodynamic Phases of Cancer Metabolism
A Spin-Glass Metabolic Hamiltonian optimized by Quantum Annealing Reveals Thermodynamic Phases of Cancer Metabolism
Sung, J.-Y.; Baek, K.; Park, I.; Bang, J.; Cheong, J.-H.
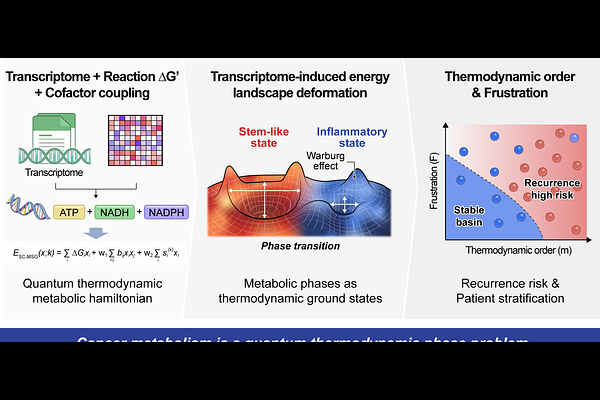
AbstractAbstract Understanding why specific metabolic states become stable in cancer has remained a fundamental challenge, as current pathway-centric frameworks lack a unifying physical principle governing global metabolic organization. We introduce the Metabolic Spin-Glass (MSG) model, which recasts cellular metabolism as a frustrated many-body system governed by a Hamiltonian that integrates reaction free energies, cofactor-mediated thermodynamic couplings, and patient-specific transcriptomic fields. The Hamiltonian is formulated as a binary optimization problem and solved using hybrid quantum annealing. Embedding gastric cancer transcriptomes (n=497) reveals that malignant phenotypes correspond to thermodynamically distinct ground states rather than isolated pathway perturbations. The Warburg effect emerges intrinsically as a thermodynamic phase transition, and stem-like tumors occupy the deepest attractor basin reflecting high energetic stability. A thermodynamic order parameter stratifies patients into prognostically distinct subtypes independently of transcriptomic classification, suggesting clinically applicable non-redundant biomarkers. This work establishes a spin-glass energy landscape framework for physically principled, patient-specific cancer metabolic stratification.