A DNA foundation model predicts osteoporosis risk genes without proximity bias
A DNA foundation model predicts osteoporosis risk genes without proximity bias
Regep, C.; Kapourani, C.-A.; Sofyali, E.; Dobrowolska, A.; Loukas, G.; Anighoro, A.; Canale, E.; Gross, T.; Licciardello, M.; Gupta, R.; Maciuca, S.; Desai, T.; Del Vecchio, A.; Field, C.; Gemayel, K.; Javer, A.; Zhang, Z.; Tsujikawa, R.; Inoue, F.; Hessel, E.; Taylor-King, J.; Whittaker, J.; Roblin, D.; McIntyre, R.; Edwards, L.
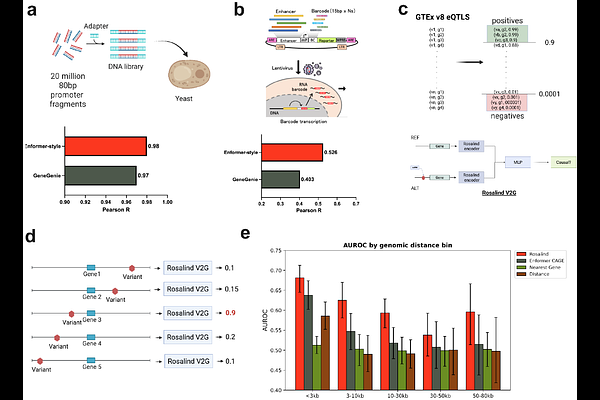
AbstractTargets supported by human genetic associations are more than twice as likely to progress from clinical development to approval. Genome-wide association studies are the largest source of genetic evidence for disease risk but linking non-coding variants to effector genes remains a significant barrier to identifying causal targets. Current gene-mapping approaches suffer from proximity bias, largely ignoring distal genes. Here we introduce Rosalind, a DNA foundation model fine-tuned on human genetic variation from GTEx, that directly predicts variant-gene regulatory relationships from sequence without relying on nearest-gene heuristics. We demonstrate Rosalind's accuracy through extensive benchmarking, apply it to multiple complex traits to establish broad utility, and provide experimental validation in osteoporosis using a translational osteoblast assay. We demonstrate that genes distal to osteoporosis risk variants were significantly more likely to alter a bone formation phenotype than nearest genes. Together, these results highlight deep learning-based regulatory models as a general and scalable framework for translating novel genetic insights to drug discovery.