Malaria control and the unexpected spread of diagnostic-resistant Plasmodium falciparum in Peru
Malaria control and the unexpected spread of diagnostic-resistant Plasmodium falciparum in Peru
Gerdes Gyuricza, I.; Fola, A. A.; Simkin, A.; Thwai, K. L.; Juliano, J. J.; Bailey, J. A.; Johri, P.; Henry, C. M.; Cabrera-Sosa, L.; Porras-Laymito, G.; Cheng, Q.; Watson, O. J.; Gamboa, D.; Valdivia, H. O.; Parr, J. B.
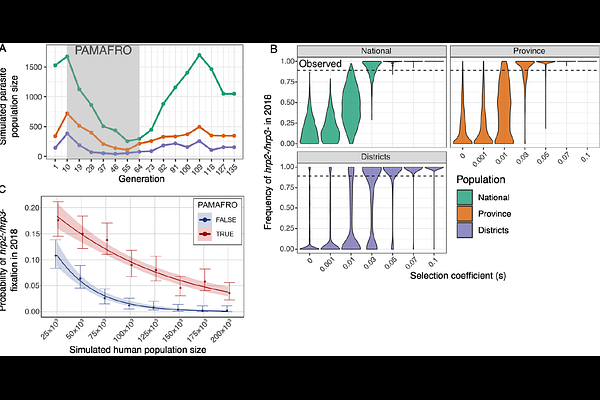
AbstractPlasmodium falciparum parasites with deletions of the histidine-rich protein 2 and 3 (hrp2 and hrp3) genes evade detection by common rapid diagnostic tests (RDTs) and pose a growing threat to malaria control. While these deletions have emerged in multiple regions globally, the evolutionary forces driving their spread remain unclear. Here, we analyze 1,215 P. falciparum samples collected between 2003 and 2018 in Loreto, Peru. This region experienced a major decline in malaria transmission following the Project for Malaria Control in Andean Border Areas (PAMAFRO) and now harbors a high proportion of hrp2/3 deleted parasites despite limited RDT use. Using molecular inversion probe (MIP) sequencing across > 2,000 genome-wide loci, we observed a marked reduction in genetic diversity, increased clonality, and fixation of parasites with deletions of both hrp2 and hrp3 genes (hrp2-/3-) over time. Identity-by-descent (IBD) analysis revealed rapid expansion of a single hrp2-/3- dominant lineage in the post-PAMAFRO period, consistent with clonal replacement after intense malaria control. Targeted sequencing of the hrp2/3 regions showed conserved deletion breakpoints across three different lineages, indicative of recombination of a common haplotype into distinct genetic backgrounds. To investigate the evolutionary forces driving the fixation of hrp2-/3- in Loreto, we simulated allele frequency trajectories under different selection coefficients. We found that fixation of hrp2-/3- due solely to genetic drift (selection coefficient s = 0) is unlikely; a selection coefficient of s [≥] 0.03 was required for fixation to occur consistently. However, our simulations also indicate that a genetic bottleneck caused by PAMAFRO increased the likelihood of fixation through drift by 4.5- to 17-fold depending on the population. These findings suggest that hrp2-/3- fixation was likely driven by a combination of demographic changes resulting from PAMAFRO and selective advantage unrelated to RDT use. Our results demonstrate how intensive malaria control efforts can reshape parasite populations and underscore the value of expanded genomic surveillance as countries move toward malaria elimination.