Distinct mechanisms of CNV formation at the human 15q13.3 locus
Distinct mechanisms of CNV formation at the human 15q13.3 locus
Höps, W.; Porubsky, D.; Yoo, D.; de Groot, M.; den Ouden, A.; Derks, R.; Hoekzema, K.; Hoischen, A.; Yntema, H. G.; Human Pangenome Reference Consortium (HPRC), ; Caro, P.; De Falco, A.; van Bon, B.; Brunetti-Pierri, N.; Schaaf, C. P.; Eichler, E.; Gilissen, C.
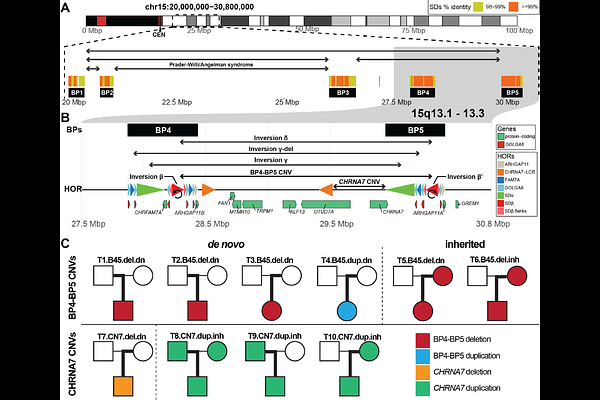
AbstractHuman chromosome 15q13.3 is a hotspot for recurrent pathogenic copy number variants (CNVs), which remain unresolved at the sequence level. We generated haplotype-resolved assemblies for 10 patient-parent trios and found that both the long (''BP4-BP5'') and short (''CHRNA7'') forms of 15q13.3 CNVs arise predominantly by non-allelic homologous recombination (NAHR) enabled by inversion polymorphisms. While most BP4-BP5 CNVs are structurally distinct, three breakpoints cluster in a 2 kbp PRDM9-enriched recombination hotspot. CHRNA7 CNVs originate from NAHR between CHRNA7-LCR repeats embedded within locus-spanning inversions and give rise to paired deletion/duplication events. Population analyses of 581 population haplotypes reveal at least 18 distinct structural haplotypes in 15q13.3 and more than 10-fold ancestry-stratification of BP4-BP5 CNV risk, where 68.4% of Europeans but only 5.1% of East Asians are predisposed. Comparison to six ape species indicates that the duplication architecture promoting instability expanded recently and is largely human-specific.