Under or Over? Tracing Complex DNA Topologies with High-Resolution Atomic Force Microscopy
Under or Over? Tracing Complex DNA Topologies with High-Resolution Atomic Force Microscopy
Gamill, M. C.; Holmes, E. P.; Provan, J. I.; Wiggins, L.; Ruskova, R.; Whittle, S.; Catley, T. E.; Main, K. H. S.; Shephard, N.; Bryant, H. E.; Racko, D.; Colloms, S. D.; Pyne, A. L. B.
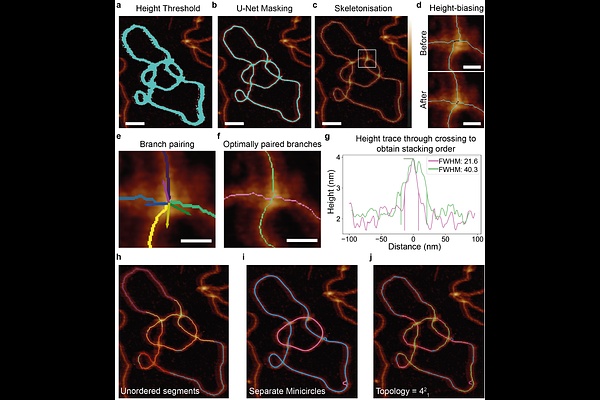
AbstractThe intricate topology of DNA plays a crucial role in the regulation of cellular processes and genome stability. Despite its significance, DNA topology is challenging to explicitly determine due to the length and conformational complexity of individual topologically constrained DNA molecules. Here, we introduce an innovative approach combining high-resolution Atomic Force Microscopy (AFM) imaging with automated computational analysis to directly determine DNA topology. Our pipeline enables high-throughput tracing of uncoated circular DNA molecules directly in aqueous conditions, enabling determination of the order of DNA crossings, i.e. which molecule passes over which. By accurately tracing the DNA path through every DNA crossing, we can explicitly determine DNA topology and precisely classify knots and catenanes. We validate our method using known catenated products of the E. coli Xer recombination system and confirm the predicted topology of knotted products from this system. Our study uncovers a recurrent depositional effect for the Xer catenanes and uncovers the origins of this effect using coarse-grained simulations, and enhances the statistical robustness. Our pipeline is applicable to other DNA/RNA-protein structures, particularly those with inherent flexibility, and opens up avenues for understanding fundamental biological processes which are regulated by or affect DNA topology.