4D Single-Cell Spatial Transcriptomics Reveals Dynamic Morphogenetic Gradients and Regenerative Domains in Planarians
4D Single-Cell Spatial Transcriptomics Reveals Dynamic Morphogenetic Gradients and Regenerative Domains in Planarians
Han, K.; Chen, Y.; Li, Y.; Guo, L.; Wang, Y.; Liu, X.; Lin, Y.; Huang, Z.; Liu, Q.; Guo, W.; Zhang, R.; Zhao, W.; Liang, L.; Wei, X.; Zhou, L.; Mao, X.; Wang, J.; Wu, W.; Pan, H.; Yang, T.; Zhang, H.; Su, X.; Liu, S.; Zhang, W.; Liu, L.; Christensen, S. T.; Fei, J.; Liu, X.; Fan, G.; Li, H.; Gu, Y.; Wang, J.; Yang, H.; Pei, G.; Xu, X.; Zeng, A.; Xu, M.
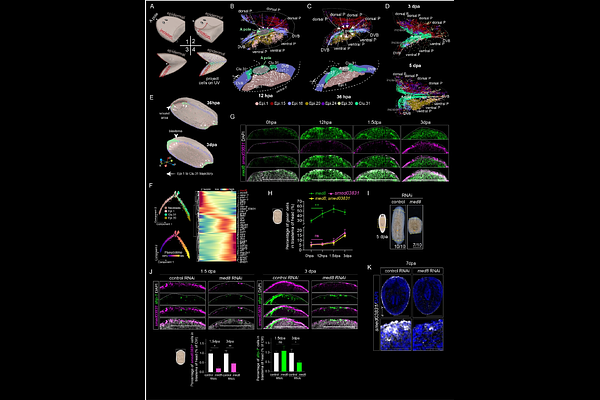
AbstractRegeneration relies on precise spatiotemporal gene expression and cellular responses to establish tissue identity and body patterning. Using high-resolution Stereo-seq (715 nm) on 353 sections from 16 whole animals at 8 regeneration timepoints, we constructed a 4D spatiotemporal transcriptomic map of planarian regeneration. Our analysis captured 36 refined cell types from 3,508,004 segmented cells, enabling genome-wide transcriptional imputation of gene expression dynamics across body axes at cellular, tissue, and organismal scales. We identified dynamic positional gradients and distinct spatially distributed cell types during regeneration, including an injury-induced Anterior Regenerative Zone (ARZ). The ARZ exhibited enriched positional signals in epidermal, muscle, and neural cells and was regulated by Mediator 8, which is crucial for polarity remodeling and blastema formation. This study provides a comprehensive spatial molecular and cellular map of regenerative processes, highlighting injury-induced spatial domains and key regulatory factors in planarian regeneration. We also provide an interactive web portal, offering a valuable resource for exploring and analyzing regeneration mechanisms in a spatiotemporal context.