Structure-guided generative design of peptides targeting the FtsQBL divisome complex inhibit Escherichia coli cell division.
Structure-guided generative design of peptides targeting the FtsQBL divisome complex inhibit Escherichia coli cell division.
Remont, P.; Liu, X.; Croci, F.; Mechaly, A.; Karimova, G.; Nguyen, M.-H.; Guijarro, J. I.; Davi, M.; Guyon, C.; Ciambur, C. B.; Agou, F.; Boucharlat, A.; Ahmed, H.; Chiaravalli, J.; Ladant, D.; Sperandio, O.
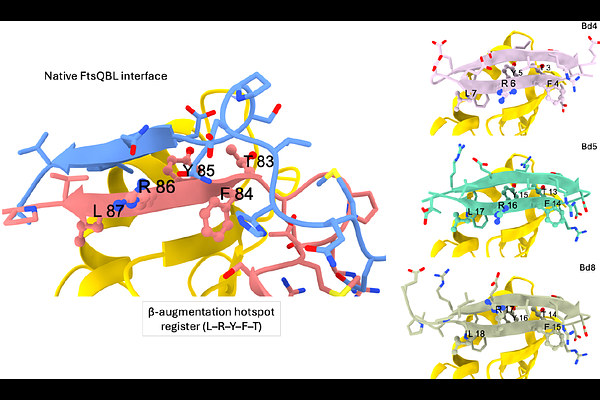
AbstractThe discovery of antibiotics targeting Gram-negative bacteria remains limited in part by the difficulty of pharmacologically modulating protein-protein interactions essential for bacterial physiology. The divisome complex formed by FtsQ, FtsB and FtsL represents an attractive but challenging target, as its assembly relies on {beta}-strand-mediated interface interactions within the bacterial periplasm. Here, we combined interpretable interface mapping using InDeep with hotspot-constrained RFdiffusion design to generate peptides targeting the FtsB-binding site of Escherichia coli FtsQ. The designed peptides mimic the native {beta}-augmentation interaction and selectively engage the FtsQ interface both in bacterial cells and in vitro. X-ray crystallography of one of these peptides in complex with FtsQ reveals that it accurately adopts the native binding geometry while introducing additional stabilizing interactions within a hydrophobic pocket. Complementary NMR analyses further show that optimized peptides adopt pre-organized {beta}-hairpin conformations in solution consistent with the bound state. Several of these peptides disrupt bacterial cell division and inhibit growth in an E. coli strain exhibiting increased outer membrane permeability. Together, these results establish a structurally validated framework in which predictive interface analysis and generative design can be combined to target cooperative protein-protein interfaces in bacteria and provide a foundation for the development of divisome-targeting antibacterial strategies.