Genetic engineering of carbon monoxide dehydrogenases produces distinct autotrophic phenotypes in Clostridium autoethanogenum
Genetic engineering of carbon monoxide dehydrogenases produces distinct autotrophic phenotypes in Clostridium autoethanogenum
Shaikh, K. M.; Reinmets, K.; Pawar, P. R.; Carneiro, C. V. G. C.; Valgepea, K.
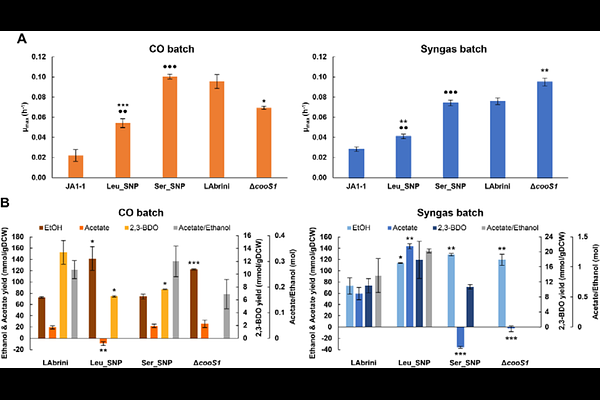
AbstractAcetogens are promising microbes for sustainable biomanufacturing but improving acetogen gas fermentation requires efficient conversion of CO and CO2 into fuels and chemicals. Carbon monoxide dehydrogenase (CODH) enzymes couple carbon fixation to energy conservation in acetogens and serve as potential regulatory modules for tuning autotrophic metabolism. Intriguingly, the model-acetogen Clostridium autoethanogenum lost its unique truncation in the bifunctional CODH (acsA), essential for autotrophy, during autotrophic adaptive laboratory evolution while obtaining superior phenotypes. Additionally, protein expression of the monofunctional CODH cooS1 is high and conditionally-regulated in C. autoethanogenum. Here, we genetically engineered CODHs in C. autoethanogenum by replacing the stop codon in acsA with leucine (strain Leu_SNP) or serine (Ser_SNP), and deleting cooS1 ({Delta}cooS1). Phenotyping in autotrophic batch and chemostat cultures revealed altered growth profiles and significant redistribution of carbon and redox flows in SNP strains, whereas {Delta}cooS1 showed moderate and condition-dependent effects. Surprisingly, structural modelling identified no conformational differences between wild-type and mutant AcsA proteins. While transcriptomics showed limited transcriptional changes in {Delta}cooS1, it suggested potential transcriptional adjustments linked to reduced robustness and altered product profile of Leu_SNP. Our results demonstrate the impact of CODHs on autotrophy and offer targets for rational engineering of acetogen cell factories.