Rapid and reliable quantification of cytosolic mRNA escape (RNASCAPE)
Rapid and reliable quantification of cytosolic mRNA escape (RNASCAPE)
Schulz, F. H.; Sorensen, E. W.; Bender, S. W.; Breuer, A.; Kyriakakis, G.; Dreisler, M. W.; Bolis, G.; Oikonomou, A.; Tsolakidis, K.; Arampatzis, S.; Nie, G.; Hatzakis, N. S.
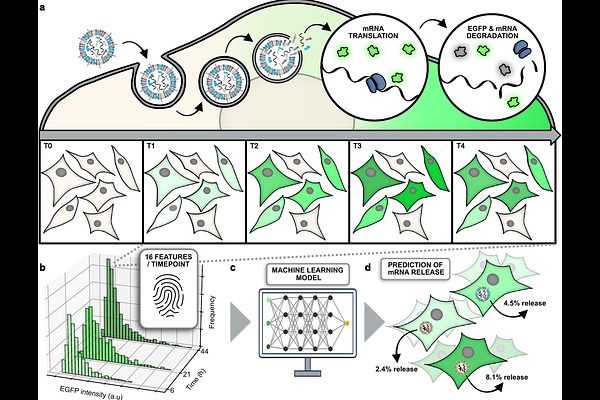
AbstractEndosomal escape of mRNA remains a critical bottleneck in oligonucleotide therapeutics, with typically less than 5% of delivered mRNA reaching the cytosol. Precise quantification of this escape is hindered by stochastic variability in uptake, release, and expression, while existing methods lack scalability and accuracy. Here we introduce RNASCAPE, a deep learning framework trained on biologically realistic simulations that estimates unlabelled mRNA cytosolic escape efficiency using only three timepoints of EGFP reporter expression and four lipid nanoparticle (LNP) meta-parameters. Benchmarking on synthetic data shows RNASCAPE achieves a mean absolute percentage accuracy of 78%. RNASCAPE accurately predicts escape efficiencies around 5-9% across diverse cell types and revealed that exchanging cholesterol to beta-sitosterol drastically drastically reduces mRNA loading and enhances twofold its functional release. By enabling robust, microscopy-agnostic quantification without requiring specialized cell lines or labeled cargo, RNASCAPE provides a scalable framework for benchmarking and rational design of LNP formulations, advancing nucleic acid therapeutic delivery.