Comparison of spatial transcriptomics technologies used for tumor cryosections
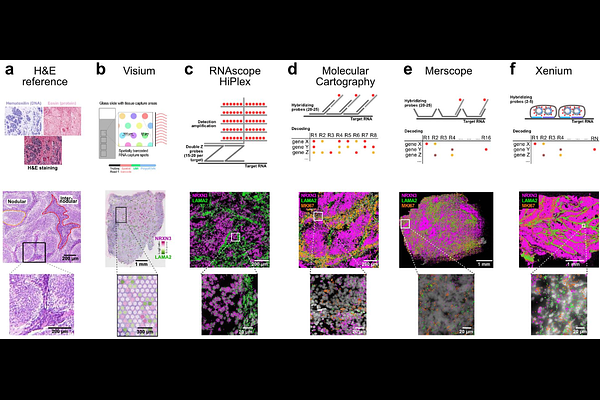
Comparison of spatial transcriptomics technologies used for tumor cryosections
Rademacher, A.; Huseynov, A.; Bortolomeazzi, M.; Wille, S. J.; Schumacher, S.; Sant, P.; Keitel, D.; Okonechnikov, K.; Ghasemi, D.; Pajtler, K.; Mallm, J.-P.; Rippe, K.
AbstractBackground: Spatial transcriptomics (ST) methods provide single cell transcriptome profiles within the endogenous cell tissue context. Thus, they are ideally suited to resolve intra-tumor heterogeneity and interactions between tumor and non-malignant cells in their microenvironment. However, different ST technologies exist and are rapidly evolving. It is frequently not clear how they address a given research question in cancer research. Here, we investigated fresh frozen tissue sections from medulloblastoma with enhanced nodularity (MBEN) using four distinct imaging-based ST methodologies (RNAscope HiPlex, Molecular Cartography, MERFISH/Merscope, and Xenium), alongside sequencing-based ST on Visium slides. Results: Our comparative case study describes critical aspects of the different workflows and identifies informative parameters to assess the experimental results. We evaluate sensitivity and specificity across platforms, show how technology dependent features affect the results and provide guidance for the experimental design. Furthermore, we demonstrate how cell segmentation can be improved and/or additional custom readouts can be integrated by reimaging of the slides after the ST experiment. Conclusions: The different ST methods come with specific features that should be considered in the method selection and experimental design. We anticipate that the insight gained in our study will facilitate successful application of ST for the analysis of fresh frozen tissue sections of solid tumors.