Cell-specific variant-to-gene mapping identifies conserved neural and glial regulators of sleep
Cell-specific variant-to-gene mapping identifies conserved neural and glial regulators of sleep
Zimmerman, A. J.; Biglari, S.; Trang, K. B.; Almeraya Del Valle, E.; Pack, A. I.; Grant, S. F.; Keene, A. C.
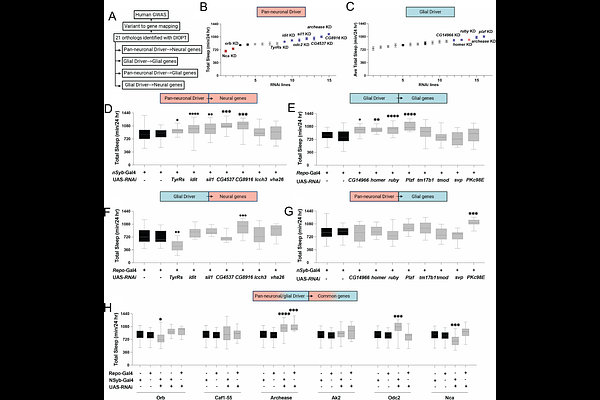
AbstractExcessive daytime sleepiness (EDS) is a heterogeneous phenotype with little known of its genetic basis. Large-scale genome-wide association studies (GWAS) have reported genomic loci associated with EDS, though since most of these are non-coding, the causal gene(s) underlying the association are not known. Additionally, the cell types in which these genes exert their effects on sleep have not been functionally explored in vivo. Here, we employed a chromatin-based variant-to-gene mapping approach to first implicate candidate effector genes at EDS GWAS loci in human-derived neural and glial cell lines. Subsequent cell type-specific RNAi knockdown of orthologous genes using neural and glial GAL4 drivers in Drosophila confirmed cell-specific regulation of sleep by these GWAS-implicated effector genes. Among these, ruby (ortholog to AP3B2), a component of the AP-3 vesicular trafficking complex emerged as a robust sleep regulator. Targeted knockdown in flies localized ruby function to astrocyte-like glia, where loss of ruby increased sleep duration. The conserved role of ruby/ ap3b2 was validated in zebrafish where CRISPR-mediated loss increased daytime sleep. Together, these findings show that physical variant-to-gene mapping predicted cell-type-specific gene function for complex sleep traits and revealed ruby/AP3B2 as a conserved glial regulator of sleep and arousal. This work provides a generalizable framework for connecting non-coding GWAS variants and their corresponding effector genes to identify novel and highly conserved regulators of sleep.