Reference genome bias in light of species-specific chromosomal reorganization and translocations
Reference genome bias in light of species-specific chromosomal reorganization and translocations
Maurstad, M. F.; Hoff, S. N. K.; Cerca, J.; Ravinet, M.; Bradbury, I. R.; Jakobsen, K. S.; Praebel, K.; Jentoft, S.
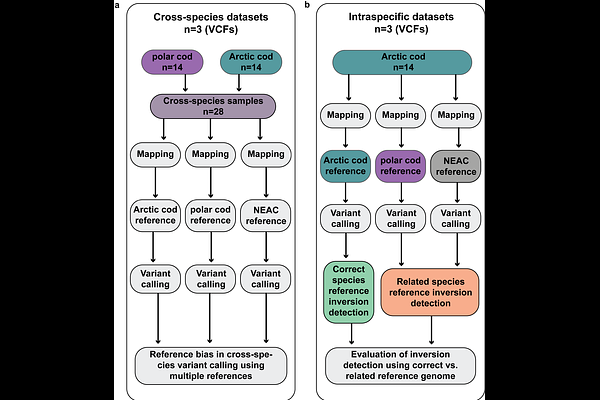
AbstractWhole-genome sequencing efforts has during the past decade unveiled the central role of genomic rearrangements - such as chromosomal inversions - in evolutionary processes, including local adaptation in a wide range of taxa. However, employment of reference genomes from distantly or even closely related species for mapping and the subsequent variant calling, can lead to errors and/or biases in the datasets generated for downstream analyses. Here, we capitalize on the recently generated chromosome-anchored genome assemblies for Arctic cod (Arctogadus glacialis), polar cod (Boreogadus saida), and Atlantic cod (Gadus morhua) to evaluate the extent and consequences of reference bias on population sequencing datasets (approx. 15-20x coverage) for both Arctic cod and polar cod. Our findings demonstrate that the choice of reference genome impacts population genetic statistics, including individual mapping depth, heterozygosity levels, and cross-species comparisons of nucleotide diversity (pi) and genetic divergence (DXY). Further, it became evident that using a more distantly related reference genome can lead to inaccurate detection and characterization of chromosomal inversions, i.e., in terms of size (length) and location (position), due to inter-chromosomal reorganizations between species. Additionally, we observe that several of the detected species-specific inversions were split into multiple genomic regions when mapped towards a heterospecific reference. Inaccurate identification of chromosomal rearrangements as well as biased population genetic measures could potentially lead to erroneous interpretation of species-specific genomic diversity, impede the resolution of local adaptation, and thus, impact predictions of their genomic potential to respond to climatic and other environmental perturbations.