Lineage-specific macrophage programs dictate metabolic suppression and stress responses associated with VBNC-like states in Listeria monocytogenes
Lineage-specific macrophage programs dictate metabolic suppression and stress responses associated with VBNC-like states in Listeria monocytogenes
Polidori, M.; Van Geest, G.; Mestre Neher, A.; Monney, C.; Bruggmann, R.; Oevermann, A.
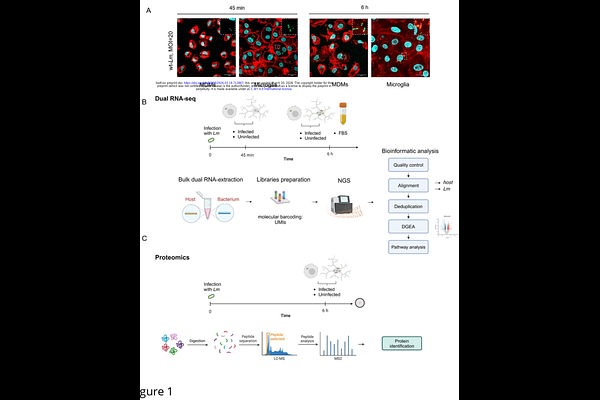
AbstractListeria monocytogenes (Lm) is a significant cause of central nervous system (CNS) infection in humans and animals, yet the mechanisms governing its intracellular lifestyle in brain-resident and infiltrating macrophages remain unclear. Using dual RNA sequencing combined with host proteomics, we map host-pathogen interactions in bovine microglia and monocyte-derived macrophages (MDMs), the two principal macrophage populations encountered by Lm in the CNS. Although microglia and MDMs share core antimicrobial programs, they differ in their metabolic and immunological states, which dictate whether Lm adopts a replicative cytosolic lifestyle or a stress-tolerant intravacuolar state. In MDMs, nutrient restriction and phagolysosomal pressure drive metabolic suppression, robust stress and SOS responses, and induction of non-coding regulatory RNAs, shifting toward a dormant, viable but non-culturable phenotype. In microglia, the nutrient-rich cytosol supports bacterial growth, marked by upregulation of nucleotide salvage and carbohydrate and lipid metabolism. Functional analyses identify the stress-related genes recA and rtcB as contributors to intracellular persistence. Together, our findings show that the fate of Lm is shaped not solely by canonical virulence genes but also by the interplay between bacterial stress adaptation and lineage-specific macrophage environments, highlighting macrophage ontogeny as a critical determinant of infection outcome.