ESPeR-seq: Extremely Sensitive and Pure, End-to-end, RNA-seq library preparation
ESPeR-seq: Extremely Sensitive and Pure, End-to-end, RNA-seq library preparation
Chen, H.-M.; Kao, J.-C.; Yang, C.-P.; Tan, C.; Lee, T.; Sugino, K.
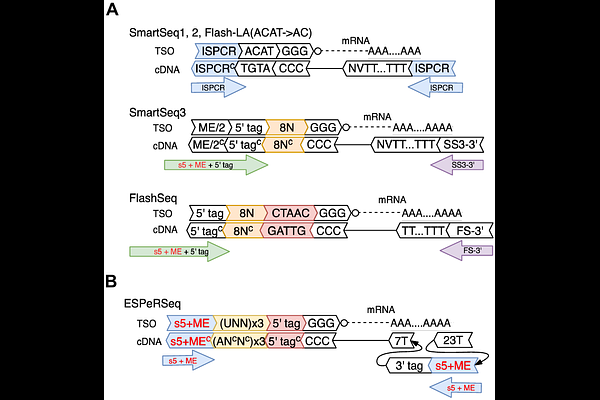
AbstractThe Smart-seq family of methods represents the gold standard for high-sensitivity, full-length single- cell RNA sequencing. Despite iterative improvements, fundamental challenges remain: the generation of non-specific PCR products that limit sensitivity, the inability to capture precise Transcription End Sites (TES), and the insidious generation of "phantom UMIs", artificial molecular barcodes created during PCR that systematically inflate molecular counts. Here, we present ESPeR-seq, a novel architecture that resolves these barriers. To enable precise, stranded TES capture, we developed an "Omega-dT" primer that bypasses synthetic poly-T tracts, restoring high-quality sequencing directly at transcript termini. To eliminate both PCR background and phantom UMIs, we implemented a biochemical "multi-lock" mechanism utilizing uracil-containing TSOs and a uracil-intolerant DNA polymerase. We validate this approach using the logQ-slope, a novel metric that sensitively diagnoses UMI fidelity. Benchmarking reveals that while state-of-the-art methods still exhibit signs of UMI inflation, ESPeR-seq strictly prevents it. Furthermore, the strandedness and precise end-delineation provided by TSO and dT reads support robust de novo gene model reconstruction, enabling the discovery of novel multi-exon genes, unannotated 3' UTR extensions, and candidate eRNAs across aggregated single-cell populations. Thus, ESPeR-seq establishes a robust framework for absolute quantitative accuracy and full-length isoform resolution.