cryoSPHERE: Single-particle heterogeneous reconstruction from cryo EM
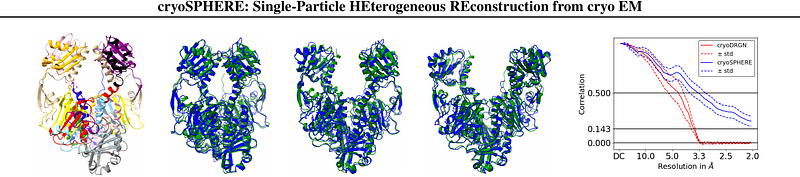
cryoSPHERE: Single-particle heterogeneous reconstruction from cryo EM
Westenhoff, S.; Grunewald, L.; Lindsten, F.; Ducrocq, G.
AbstractThe three-dimensional structure of a protein plays a key role in determining its function. Methods like AlphaFold have revolutionized protein structure prediction based only on the amino-acid sequence. However, proteins often appear in multiple different conformations, and it is highly relevant to resolve the full conformational distribution. Single-particle cryo-electron microscopy (cryo EM) is a powerful tool for capturing a large number of images of a given protein, frequently in different conformations (referred to as particles). The images are, however, very noisy projections of the protein, and traditional methods for cryo EM reconstruction are limited to recovering a single, or a few, conformations. In this paper, we introduce cryoSPHERE, a deep learning method that takes as input a nominal protein structure, e.g. from AlphaFold, learns how to divide it into segments, and how to move these as approximately rigid bodies to fit the different conformations present in the cryo EM dataset. This formulation is shown to provide enough constraints to recover meaningful reconstructions of single protein structures. This is illustrated in three examples where we show consistent improvements over the current state-of-the-art for heterogeneous reconstruction.


