Experimental Data Driven AI Framework for Flexible Protein Conformational Reconstruction
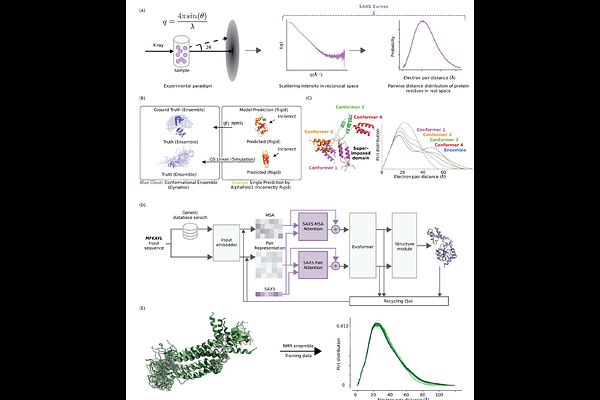
Experimental Data Driven AI Framework for Flexible Protein Conformational Reconstruction
Yu, F.; Prince, S.; Tritt, A.; Pande, K.; Hura, G. L.; Ruebel, O.; Tsutakawa, S. E.
AbstractDeep learning has revolutionized structural biology by prediction with near experimental accuracy static protein folds from amino acid sequence alone. However, proteins function as dynamic ensembles of protein conformation states, and current sequence-only models often fail to capture the specific conformational states and heterogeneity dictated by cellular environments or ligand binding. While recent generative models can sample broad conformational landscapes, they remain unconstrained by physical reality, often hallucinating plausible but experimentally invalid states. Here, we present AlphaSAXS, an end-to-end framework that constrains artificial intelligence (AI) inference using Small Angle X-ray Scattering (SAXS) experimental solution scattering data. By integrating real-space pair distance distributions (P(r)) directly into the AlphaFold architecture, AlphaSAXS effectively steers the structural hypothesis toward the experimentally observed structures. We demonstrate that AlphaSAXS resolves documented failure modes of sequence-only models in Apo-Holo transitions, successfully distinguishing between states with identical sequences but distinct scattering profiles. Furthermore, we introduce a hybrid inference protocol that couples deep learning with biophysical hydration modeling, enabling the reconstruction of solution state protein ensembles compatible with experimental data. This work establishes a paradigm for experimentally guided AI, bridging the gap between probabilistic sampling and biophysical measurement.