Epigenomic profiling of cerebrospinal fluid cells identifies immune regulatory alterations and implicates protocadherins in multiple sclerosis
Epigenomic profiling of cerebrospinal fluid cells identifies immune regulatory alterations and implicates protocadherins in multiple sclerosis
Han, Y.; Zheleznyakova, G. Y.; Sorini, C.; Pahlevan Kakhki, M.; Ruffin, N.; Liang, H.; Hallen, N.; Rao Prakash, C.; Beckers, V.; Ivanova, E.; Khademi, M.; Karlsson, M. C. I.; Piehl, F.; Olsson, T.; Kelsey, G.; Kular, L.; Needhamsen, M.; Jagodic, M.
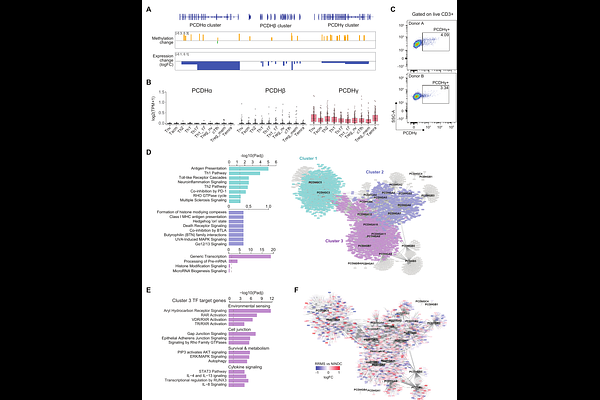
AbstractMultiple sclerosis (MS) is a chronic inflammatory disease of the central nervous system (CNS), where DNA methylation may play a role by connecting genetic and environmental risk factors. We performed whole-genome DNA methylation profiling of cerebrospinal fluid (CSF) cells from relapsing-remitting MS patients and matched controls, identifying 2,710 differentially methylated positions (DMPs) and 4,330 regions (DMRs). These changes were enriched in immune signaling, adhesion and migration processes, and were accompanied by corresponding RNA expression changes. MS-associated methylation changes enriched in the cohesin chromatin regulation pathway mapped to enhancers of T helper 17 (Th17) cells, whereas in other T cell types they were mapping to bivalent enhancers and repressed chromatin. Notably, this pathway comprised multiple Protocadherin (PCDH) genes, typically expressed in neuronal cells, that displayed consistent methylation and expression changes in CSF cells. Expression of shared intracellular domain of PCDH{gamma} cluster proteins was confirmed in peripheral blood T cells by flow cytometry as well as expression of PCDH{gamma} cluster genes in memory CD4+ T cell subsets. Moreover, co-expression analysis suggests a role of PCDH genes in aryl hydrocarbon receptor (AHR) signaling. In summary, DNA methylation changes in CSF resident cells reflect dysregulated T cell activation and migration in MS and suggest a novel role of protocadherin molecules in MS pathogenesis.