Nucleotide-dependent Structural Selection Governs c-Src Phosphorylation of Oncogenic KRas4B-G12D
Nucleotide-dependent Structural Selection Governs c-Src Phosphorylation of Oncogenic KRas4B-G12D
Lu, H.; Xu, H.; Marti, J.; Ma, B.; FARAUDO, J.
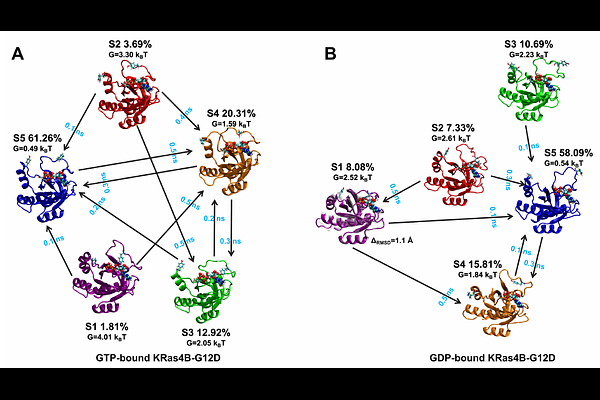
AbstractKirsten rat sarcoma viral oncogene homologue (KRas) is a frequently mutated oncogene in many types of cancer and its disregulation is frequently linked to cancer progression. Phosphorylation of KRas by tyrosine kynase c-Src induces conformational changes that weakens KRas interaction with downstream effectors. It is known that c-Src preferentially recognizes and phosphorylates the GTP-bound form of KRas over its GDP-bound counterpart, but the molecular mechanism of this nucleotide-dependent selectivity remains unknown. Here, we identify the molecular mechanism underlying the c-Src recognition of the GTP-loaded state of KRas by combining extensive all-atom molecular dynamics simulations with Markov State Models analysis. In our simulations, we considered a particularly relevant KRas mutant (KRas4B-G12D, which is the most prevalent in humans) both in GDP and GTP loaded states. Our results reveal that c-Src preferentially engages the dominant macrostates of GTP-loaded KRas4B-G12D, whereas productive interactions with the GDP-bound form occur primarily through sparsely populated conformations. We identify two Src interaction regions (residues 340-359 and 453-473) that form state-dependent contacts selectively stabilizing phosphorylation-competent KRas conformations, and they are minimally engaged in other macrostates of KRas. These highly populated conformational states of GTP-loaded Kras selectively recognized by c-Src constitute privileged targets for the rational design of peptide-based or small-molecule inhibitors that specifically target active KRas4B-G12D while sparing the inactive GDP-bound form.