Female iPSC X-chromosome inactivation (XCI) erosion and its transcriptomic effects during CRISPR gene editing and neural differentiation
Female iPSC X-chromosome inactivation (XCI) erosion and its transcriptomic effects during CRISPR gene editing and neural differentiation
Thapa, C.; Oh, E. K.; Sirkin, D.; Lahey, J.; Diaz de Leon Guerrerro, S.; McCarroll, A.; Gowda, P.; Zhang, H.; Barishman, A.; Peyton, L.; Zhang, S.; Pollak, R. M.; Hart, R. P.; Pato, C. N.; Kreimer, A.; Mulle, J. G.; Sanders, A. R.; Pang, Z.; Duan, J.
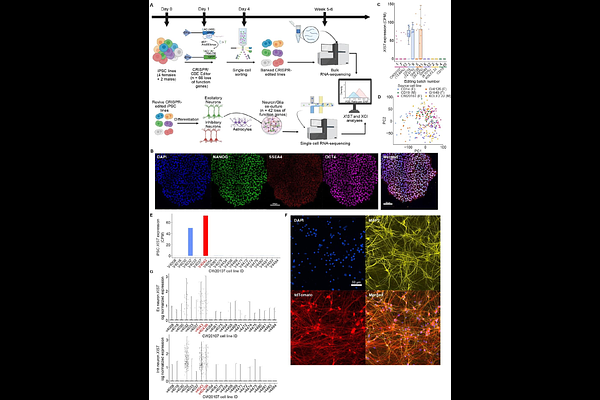
AbstractHuman induced pluripotent stem cells (hiPSC) and iPSC-differentiated neural cells, in combination with CRISPR editing, are commonly used for studying neurodevelopmental and other brain disorders. Female iPSCs undergo random X-chromosome inactivation (XCI) via epigenetic silencing by noncoding X inactive specific transcript (XIST). It is known that female iPSCs may lose XIST expression, leading to XCI erosion that affects both X-linked and autosomal gene expression. However, the effects of CRSIPR editing and neural differentiation on XCI erosion in iPSC-derived neurons and how this may confound a real-world transcriptomic analysis of differentially expressed genes (DEGs) are poorly understood. Here, leveraging bulk RNA-seq of hundreds of CRISPR-edited female iPSC lines from four donor lines for 66 genes and single-cell RNA-seq of iPSC-derived neurons of a subset of 42 edited genes, we investigated the effects of XCI erosion during CRISPR editing and in iPSC-derived neurons. We found that XCI erosion was variable in CRISPR-edited female iPSCs and largely preserved in iPSC-derived neurons. Like in iPSCs, XIST in neurons predominately influenced the expression of X-linked genes; however, its effect on autosomal genes was more pronounced in single neurons. Mechanistically, XIST epigenetically causes allelic imbalance of both X-linked and autosomal genes, with the former showing stronger allele-specific expression (ASE) bias. Notably, XIST-induced ASE bias exhibited a conserved positional pattern at loci affecting neurodevelopmental genes across different female lines and cell types. Finally, we demonstrated a confounding effect of XCI erosion on DEG analyses in iPSC-derived neurons. These results have significant implications in hiPSC modeling of neurodevelopmental and other brain disorders.