A functional genetic landscape of antibiotic sensitivity across the pneumococcal pangenome reveals conserved and lineage-specific vulnerabilities
A functional genetic landscape of antibiotic sensitivity across the pneumococcal pangenome reveals conserved and lineage-specific vulnerabilities
Sewgoolam, B.; Janssen, A. B.; Martin, L. S.; Rengifo-Gonzalez, M.; de Bakker, V.; Rozendal, B.; Cremers, A. J.; Veening, J.-W.
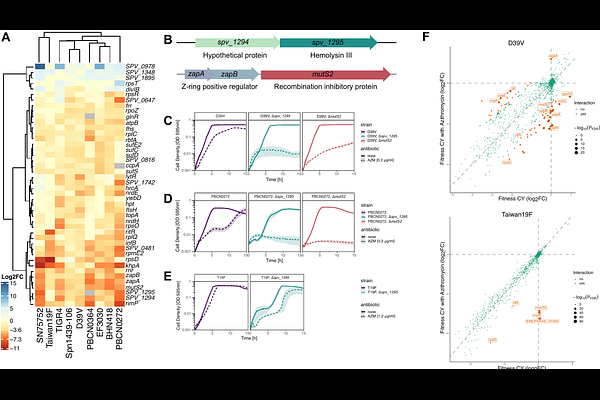
AbstractThe large pangenome of Streptococcus pneumoniae enables this opportunistic pathogen to adapt and evade antibiotic treatment. Effective treatment of pneumococcal infections requires a better understanding of the genes that modulate susceptibility to antibiotics across the pangenome. Using CRISPRi-seq, we identified genes that contribute to antibiotic sensitivity against a panel of clinically relevant antibiotics across nine pneumococcal strains with diverse resistance profiles, serotypes, and lineages. The here-generated chemical-genetics atlas revealed distinct genome-wide signatures of antibiotic stress that were specific to the antibiotic mode of action and showed both strain-specific and conserved signatures. This allowed us to identify conserved genes involved in antibiotic vulnerability and assign functions to previously uncharacterized genes. For instance, deletion of mutS2, which may act as a ribosome collision sensor and spv_1295, a conserved gene of unknown function, resulted in increased sensitivity to the macrolide azithromycin across strains, including a macrolide resistant strain, and could be potential targets for global sensitizing therapies. This work establishes a pangenome-wide framework for understanding antibiotic stress responses in S. pneumoniae, providing a foundation for the rational development of therapies that exploit conserved and strain-specific vulnerabilities.