In silico analysis reveals the structural basis of TomEP specificity, a tomato extensin peroxidase
In silico analysis reveals the structural basis of TomEP specificity, a tomato extensin peroxidase
Hussain, Z.; Sharma, S.; Faik, A.; Held, M. A.
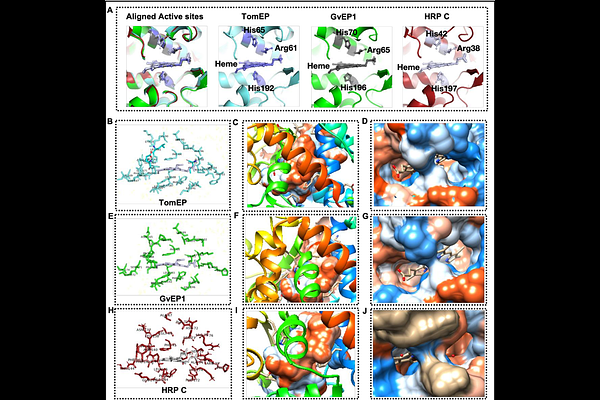
AbstractBackground Extensin peroxidases (EPs) are class III plant peroxidases and are responsible for intermolecular covalent crosslinking of extensin (EXT) monomers to create scaffolds within plant cell walls. The formation of these scaffolds impacts plant development, mechanical wounding, and response to pathogen attacks. Therefore, elucidating the molecular mechanism controlling covalent crosslinking of EXT monomers is crucial for understanding cell wall deposition and potentially improving plant growth and adaptation. The focus of this work is to use in silico analysis to determine the structural characteristics of an EP from tomato (TomEP) to elucidate its specificity for crosslinking of EXT monomers. Results In this study the two-dimensional (2D) and three-dimensional (3D) structures of TomEP were determined using several advanced bioinformatics tools and compared to two other peroxidases: GvEP1 (a known EP) and HRP-C (having a low affinity for EXT substrates). The results revealed that TomEP is a stable and hydrophilic protein with high thermal stability. The heme binding pockets of TomEP and GvEP1 have more hydrophobic residues and larger volume and pocket area compared to HRP-C. Molecular docking at the active site, which includes a heme heteroatom, showed that the ligands consisting of the hydrophobic Tyrosine-X-Tyrosine [-Y-X-Y-] motifs (i.e., [-Y-K-Y-], [-Y-V-Y-], and [-Y-Y-Y-] found in EXTs, and their derivatives, Isodityrosine (IDT), Pulcherosine (Pul), Di-Isodytirosine (diIDT), bind perfectly to the active site of TomEP with dominant interactions of Val54, Ser94, Ala96 and Phe196 residues. Pulcherosine had the highest binding affinity, while [-Y-K-Y-] showed the lowest binding affinity. Molecular dynamics simulations showed that [-Y-X-Y-] motifs (and the derivative substrate ligands) remain bound to the active site of TomEP throughout the 100 ns long simulation. Furthermore, the binding of these substrates stabilized the protein structure. Conclusion These results may explain why TomEP is particularly well-suited for EXT crosslinking and will have significant implications on biochemistry, biotechnology, and the potential use of these EPs in crops improvement.