Spatial confinement reshapes the folding of an ion-stabilized DNA with three-way junction
Spatial confinement reshapes the folding of an ion-stabilized DNA with three-way junction
Wang, X.; Shi, Y.-Z.
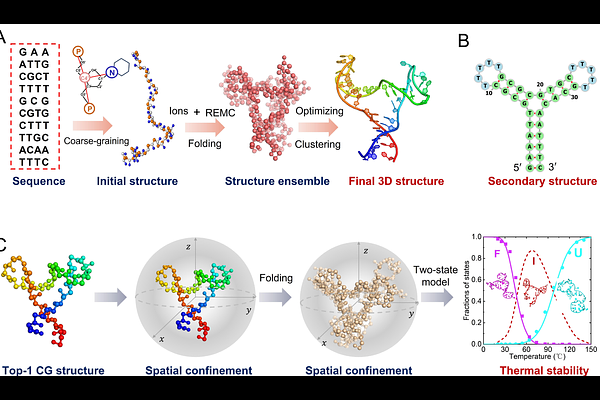
AbstractWithin the densely packed cellular milieu, the structural dynamics and stability of complex DNA architectures are governed by an intricate interplay between spatial confinement and ionic environment. Here we employ our coarse-grained model, DNAfold2, to dissect the mechanistic principles underlying how nanoscale confinement reshapes the folding landscape of a DNA with three-way junction across a broad range of ionic conditions. Our simulations demonstrate that spatial confinement acts as a structural selector, preferentially stabilizing compact, well-defined junction topologies while significantly reducing conformational flexibility in response to ionic strength variations. Through rigorous analysis of structural state populations, we reveal that this behavior results from the entropic exclusion of extended intermediates and unfolded states, effectively reshaping the DNA stability under spatial confinement. Notably, spatial confinement induces a fundamental reshape of the unfolding pathway, suppressing heterogeneous intermediate ensembles and enforcing a highly cooperative transition. These findings establish a unified mechanistic framework for understanding the biophysics of nucleic acid in physiologically crowded environments, with significant implications for de novo design of biomolecular nanomaterials and mechanistic regulation of DNA-processing machinery in vivo.