Quantitative 3D cytoarchitecture of human brain organoids using light-sheet microscopy
Quantitative 3D cytoarchitecture of human brain organoids using light-sheet microscopy
Vinchure, O. S.; Job, A. V.; Alonso-Olivares, H.; Alkuraya, F. S.; Gabriel, E.; Gopalakrishnan, J.
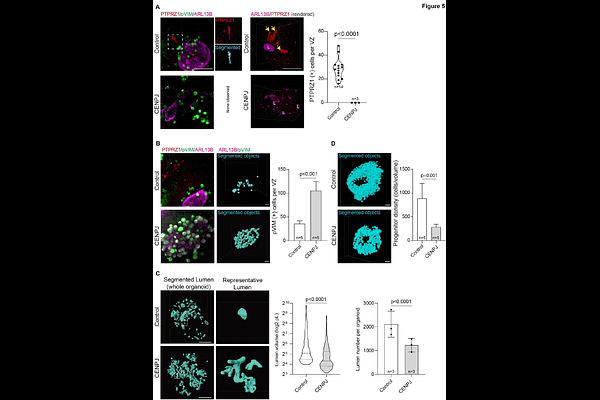
AbstractHuman brain organoids are valuable models for studying human brain development and disorders. Although organoids enable multi-omics analyses, understanding developmental mechanisms requires precise three-dimensional mapping of cell types and their dynamics within their cytoarchitecture. Traditional thin-section immunohistochemistry and imaging provide insights but disrupt spatial information. Light-sheet microscopy allows volumetric imaging of intact organoids, but methods and analytical algorithms remain technically challenging. Here, we introduce LUCID-org, an optimized and reproducible technique for Light-sheet imaging of Unsectioned, Cleared, Immunolabeled, and Depth-resolved human brain organoids. Coupled with machine-learning-based quantitative analysis, LUCID-org enables unbiased 3D characterization of cell-type composition and cytoarchitecture. We validated this approach using brain organoids derived from a patient with CENPJ-mutated microcephaly, revealing quantifiable defects in the ventricular zone (VZ), progenitor density, dynamics, cell-type variability, VZ lumen volume, and neuronal distribution compared with healthy controls. The LUCID-org pipeline is straightforward, taking approximately one week, significantly cheaper, non-toxic, and compatible with standard light-sheet microscopes. Therefore, the LUCID-org method could serve as a benchmark for comprehensive 3D analysis of human brain organoids.