Critical amino acid residues in the N-terminal domain of NADPH-dependent assimilatory sulfite reductase flavoprotein mediate octameric assembly
Critical amino acid residues in the N-terminal domain of NADPH-dependent assimilatory sulfite reductase flavoprotein mediate octameric assembly
Walia, N.; Pedrete, T.; Ahmadizadeh, F.; Rahman, E.; Garg, Y.; Washburn, B.; Pye, C.; Liu, F.; Randolph, P. S.; Weiss, K. L.; Nagy, G.; Bleiholder, C.; Stroupe, M. E.
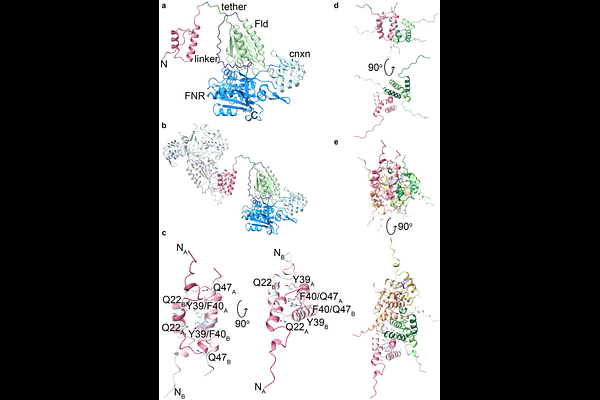
AbstractHow large, flexible enzymes assemble into defined oligomeric architectures remains a central question in biology. NADPH-dependent assimilatory sulfite reductase (SiR) forms a heterododecamer built on an octameric flavoprotein (SiRFP) core, yet the molecular basis for this assembly has been unresolved because of its disordered N-terminus. Here, we use ion mobility mass spectrometry, small-angle neutron scattering, and mutagenesis to define the mechanism of SiRFP oligomerization. We show that SiRFP forms a discrete, stable octamer in solution. We also report that its N-terminal 52-residue segment is necessary and sufficient to mediate assembly, also mediating oligomerization when fused to a heterologous protein. Structure-guided mutagenesis identifies four residues (Gln22, Tyr39, Phe40, and Gln47) whose substitution disrupts the octamer, producing concentration-dependent lower-order species while retaining catalytic activity. These findings define the determinants of SiRFP assembly with broader implications for engineering homomeric protein complexes.