Emergence of a multidrug-resistant Salmonella enterica serovar Amager lineage carrying the blaCTX-M-65-positive pESI megaplasmid
Emergence of a multidrug-resistant Salmonella enterica serovar Amager lineage carrying the blaCTX-M-65-positive pESI megaplasmid
Miranda-Riveros, J.; Tichy-Navarro, D.; Navarrete, M. J.; Reyes-Jara, A.; Toro, M.; Ugalde, J. A.; Moreno-Switt, A. I.; Pina-Iturbe, A.
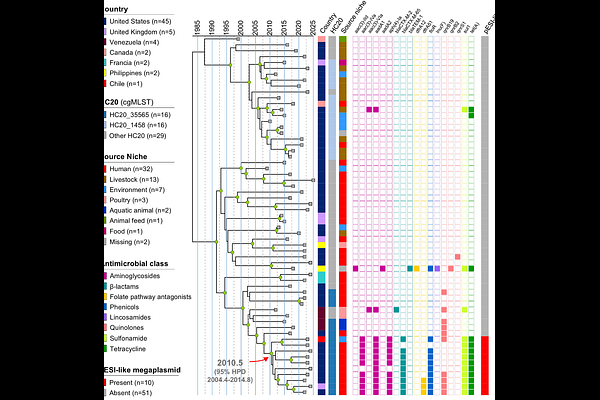
AbstractThe spread of extended-spectrum {beta}-lactamase (ESBL)-producing and fluoroquinolone-resistant Salmonella pose a global public health challenge in addition to the high burden of infections associated with this foodborne pathogen. In this study we aimed to characterize a multidrug-resistant strain of Salmonella serovar Amager isolated from a Chilean river in October 2023. Antimicrobial susceptibility testing revealed a resistance phenotype against multiple antibiotic families, including fluoroquinolones and {beta}-lactams, showing ESBL production. Hybrid genome sequencing allowed the identification of a 311,303 bp plasmid carrying the aadA1, aph(4)-Ia, aac(3)-IVa, floR, sul1, tet(A), and blaCTX-M-65 genes, sharing 99.98% sequence identity with the Salmonella Infantis pESI-like megaplasmid. In addition, the qnrB19 gene was found in a {approx}2.7 kbp plasmid of widespread distribution. Population structure and temporal phylogenetic analysis at the global scale revealed the emergence of a Salmonella Amager lineage from the HC20_35565 cluster, carrying the Salmonella Infantis blaCTX-M-65-positive pESI-like megaplasmid and causing human infections in the United States and the United Kingdom. Our work describes the emergence of a Salmonella lineage with resistance against first-line antibiotics used for treating severe infections, underscoring the relevance of environmental surveillance as a means for detecting emergent pathogens and anticipating human infections.