Mechanical signatures of nucleic acid knot topology
Mechanical signatures of nucleic acid knot topology
Bakker, D. t. R.; Yang, M.; Li, I. T. S.
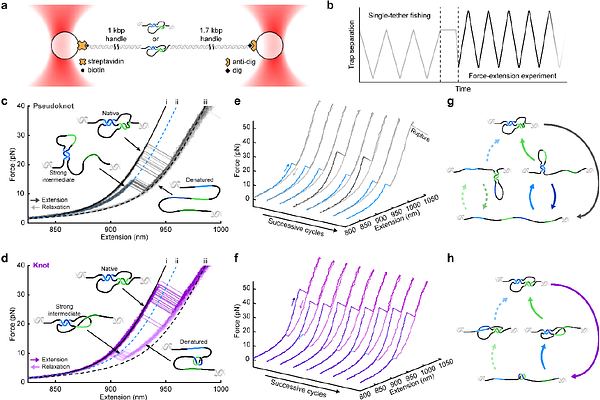
AbstractMolecular knots represent a fundamental form of polymer topology, yet their mechanical behaviour in nucleic acids remains largely unexplored. Here, we engineer a single-stranded DNA sequence that can fold into either a knot or a pseudoknot while maintaining identical base-pairing interactions. Using single-molecule force spectroscopy with optical tweezers, we show that molecular topology alone produces distinct mechanical behaviour. Knotted ssDNA exhibits three characteristic signatures relative to the pseudoknot: higher unfolding forces, shorter unfolding extensions, and faster refolding kinetics. These features arise from the topological constraint imposed by strand threading and together provide a mechanical fingerprint that distinguishes knotted from unknotted nucleic acid structures. By analyzing the denatured state under tension, we further show that the knot tightens as force increases, entering a tight-knot regime in which the molecule can be described as a compact knot core in series with a stretched single strand. The retained contour length reveals that the tight trefoil knot contains approximately ten nucleotides at forces approaching 40 pN. These results establish mechanical signatures as a means of identifying nucleic acid topology and provide quantitative insight into the nanomechanics of molecular knots.