Novel Prion Protein Gene (PRNP) Variants in Wild Montana Mule Deer
Novel Prion Protein Gene (PRNP) Variants in Wild Montana Mule Deer
Seerley, A. L.; Rothfuss, M. T.; Gray, B. M.; Sebogo, M. A.; Manakelew, B. A.; Pounder, J. I.; Bowler, B. E.; Leavens, M. J.; Grindeland Panter, A. L.
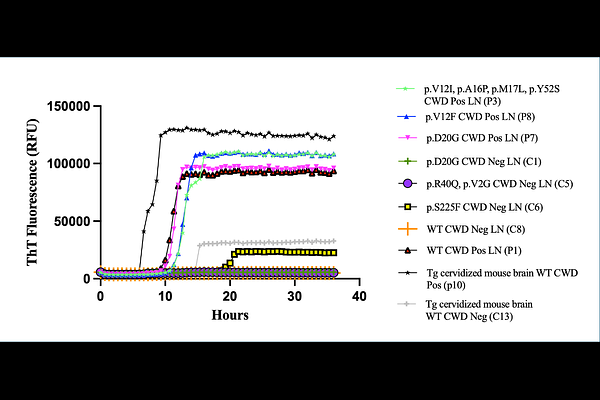
AbstractChronic Wasting Disease (CWD) is a transmissible spongiform encephalopathy (TSE) of cervids (elk, deer, moose, and reindeer) that is increasing in prevalence and expanding to new geographical areas. TSEs, commonly referred to as prion diseases, are fatal neurodegenerative diseases that occur in a variety of mammals, including humans, and typically exhibit species-specific characteristics. This study reports the sequencing of the prion protein gene (PRNP) in retropharyngeal lymph node samples from 358 Montana mule deer (Odocoileus hemionus) and the identification of 36 PRNP genetic variants, many of which have not been reported previously. Further investigations tracked spatiotemporal characteristics of variants to hunting districts, year of harvest, and CWD status. PRNP polymorphisms V12F, D20G, R40Q, and S225F were examined with EmCAST computational predictions to determine the relationship between sequence and structural variations providing further insights into mechanisms affecting CWD outcomes. EmCAST predictions suggest the novel variant V12F phenotype is attributable to functional changes such as altered protein-protein interactions that might be linked to the CWD positive status of the samples. Notably, the analysis of S225F by EmCAST predicted that S225F is a neutral mutation for folded PrP and incompatible with fibril PrP, suggesting a potential structural mechanism for why this previously known variant may provide protection against CWD based on reduced fibril PrP formation. The CWD-positive samples harboring PRNP variants were examined with the prion RT-QuIC assay, including the novel variant V12F, which resulted in prion seeding activity.