Unpaired TCRα + TCRβ sequencing is sufficient for training machine learning TCR-epitope recognition predictors
Unpaired TCRα + TCRβ sequencing is sufficient for training machine learning TCR-epitope recognition predictors
Shah, A.; Genolet, R.; Auger, A.; Moreno, D. L.; Liu, Y.; Croce, G.; Racle, J.; Harari, A.; Gfeller, D.
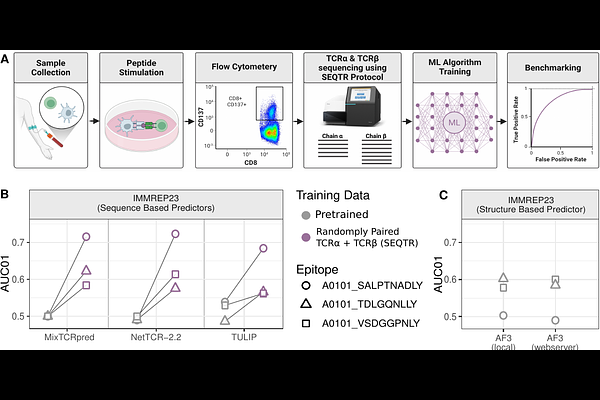
AbstractT-cell recognition of infected and malignant cells is elicited by the binding of heterodimeric T-Cell Receptors (TCRs) to epitopes and both the TCR and the TCR{beta} chains play a key role in these interactions. Machine learning tools trained on databases of TCRs recognizing diverse epitopes are useful for identifying epitope-specific TCRs in large TCR repertoire datasets. However, collecting paired TCR{beta} sequences to train such tools is associated with significant sequencing costs. Here we demonstrate that unpaired TCR + TCR{beta} sequencing of epitope-specific T cells can be used for training TCR-epitope recognition predictors at a much reduced cost compared to standard single-cell TCR sequencing protocols and with no impact on prediction accuracy. Applying this approach to some unseen epitopes used in the IMMREP community benchmark demonstrates improved accuracy compared to both existing machine learning models and AlphaFold3-based predictions.