TS2CG as a membrane builder
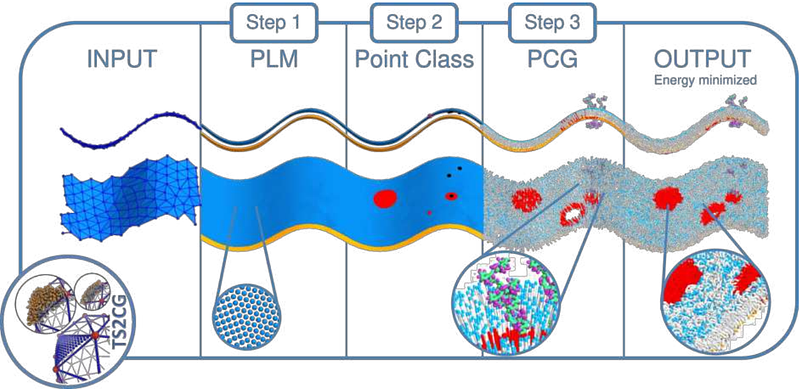
TS2CG as a membrane builder
Schuhmann, F.; Stevens, J. A.; Rahmani, N.; Lindahl, I.; Brown, C. M.; Brasnett, C.; Anastasiou, D.; Bravo Vidal, A.; Geiger, B. J.; Marrink, S.-J.; Pezeshkian, W.
AbstractMolecular dynamics (MD) simulations excel at capturing biological processes at the molecular scale but rely on a well-defined initial structure. As MD simulations now extend to whole-cell-level modeling, new tools are needed to efficiently build initial structures. Here, we introduce TS2CG version 2, designed to construct coarse-grained membrane structures with any desired shape and lateral organization. This version enables precise placement of lipids and proteins based on curvature preference, facilitating the creation of large, near-equilibrium membranes. Additional features include controlled pore generation and the placement of specific lipids at membrane edges for stabilization. Moreover, a Python interface allows users to extend functionality while maintaining the high performance of the C++ core. To demonstrate its capabilities, we showcase challenging simulations, including a Moebius strip membrane, a vesicle with lipid domain as continental plates (Martini globe), and entire mitochondrial membranes exhibiting lipid heterogeneity due to curvature, along with a comprehensive set of tutorials.