Cross hybridization Inference for Phylogenetic Resolution (CIPHR)-FISH enables microbiome imaging with strain level taxonomic resolution
Cross hybridization Inference for Phylogenetic Resolution (CIPHR)-FISH enables microbiome imaging with strain level taxonomic resolution
Adade, E. E.; Wang, R.; Henneberry, C. M.; Lemus, A. A.; Stevick, R. J.; Perez-Pascual, D.; Audrain, B.; Orsino, A. J.; Farnsworth, D. R.; Ghigo, J.-M.; Valm, A. M.
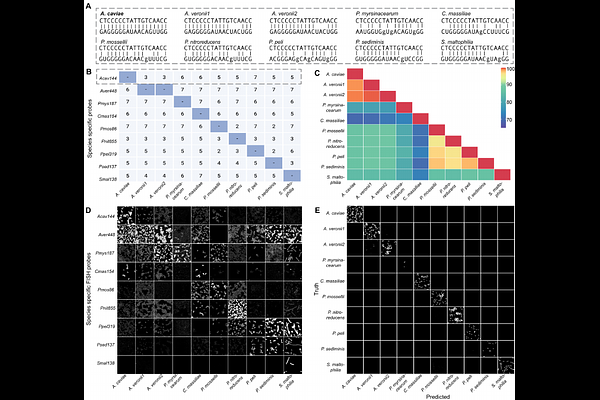
AbstractThe spatial organization of microbial communities is a critical determinant of host-microbe interactions, yet species-level mapping remains challenging due to high 16S rRNA sequence homology and spectral crosstalk in multiplexed fluorescence in situ hybridization (FISH). To address this challenge, we developed Cross-hybridization Inference for Phylogenetic Resolution (CIPHR)-FISH, a pipeline that integrates strategic probe design with supervised machine learning. CIPHR-FISH transforms probe cross-hybridization and spectral overlap, traditionally viewed as experimental noise, into informative molecular signatures. Using a gnotobiotic zebrafish model colonized with a defined mix of 10 zebrafish bacterial strains, we trained a support vector machine (SVM) on empirical hybridization patterns from pure bacterial cultures. CIPHR-FISH achieved 99.2 % macro-averaged accuracy, significantly outperforming standard linear unmixing (62.5 %), and successfully discriminated strains with 99.7% sequence homology. Applying this tool to gnotobiotic zebrafish larvae revealed distinct biogeographies: the intestinal bulb hosted highly structured, multi-layered polymicrobial aggregates, while the skin exhibited sparse, uniformly dispersed individual bacterial cells. Notably, we observed significant inter-individual variation in spatial community structure that was obscured by traditional bulk 16S rRNA sequencing. CIPHR-FISH provides a robust, scalable framework for high-resolution spatial biology by converting the limitations of molecular labeling into a rich data source for taxonomic classification. This approach enables the quantification of micro-scale ecological and stochastic forces that shape the microbiome across hosts.