Single-cell CRISPR activation screens in primary B cells discover gene regulatory mechanisms for hundreds of autoimmune risk loci.
Single-cell CRISPR activation screens in primary B cells discover gene regulatory mechanisms for hundreds of autoimmune risk loci.
Kriachkov, V.; Ching, J. W. H.; Lancaster, J.; Vespasiani, D.; Denny, N.; Hamley, J. C.; Gubbels, L.; Bandala Sanchez, E.; Neeland, M.; Levi, E.; Davies, K.; Shanthikumar, S.; Shevchenko, G.; Bryant, V. L.; Hodson, D. J.; Davies, J. O. J.; King, H. W.
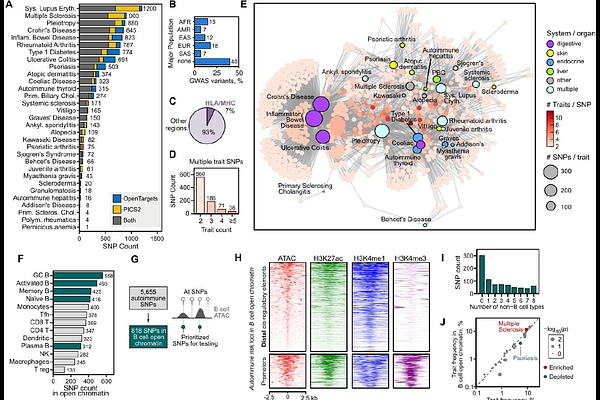
AbstractGenome-wide association studies (GWAS) have discovered thousands of genetic variants linked to autoimmune disease, and yet the molecular pathways underlying autoimmunity have remained elusive. A key challenge is that >90% of identified GWAS risk loci are in non-coding genomic regions making it difficult to predict their relevance to disease. Here, we have curated fine-mapped non-coding risk variants from over 30 different autoimmune traits including common conditions such as systemic lupus erythematosus (SLE), Crohn's disease, and multiple sclerosis, and reveal shared genetic signatures between diverse autoimmune diseases. We subsequently performed a high-throughput single-cell multi-omic CRISPR activation screen targeting 763 autoimmune risk loci in primary human B cells (a highly relevant cell type to autoimmune diseases) and discover 524 cis-regulatory target gene effects for 378 risk loci, with many risk loci regulating multiple gene targets. This Single Cell Analysis of Non-coding Distal Autoimmune Loci (SCANDAL) provides a powerful experimental resource linking non-coding risk loci to many disease-relevant genes, including lowly-expressed cytokines and transcription factors for which perturbation effects can be difficult to quantify with other CRISPR-based strategies. We reveal how increased transcriptional activity at one non-coding risk locus can drive transcription at other risk loci within the same regulatory landscape that may be relevant to understand genetic pleiotropy of autoimmune diseases. Finally, we quantified allele-specific effects on target gene expression with massive parallel reporter assays and prime editing to discover a gain-of-function variant associated with SLE that controls expression of the transcription factor REL/cREL which subsequently binds dozens of risk loci and target genes associated with different autoimmune diseases. Our study provides a valuable resource linking non-coding risk loci with their cis-regulatory target genes and advances our understanding of the shared genetic networks and mechanisms involved in autoimmunity.