Metabolic specialization structures gut bacterial niches and drives colorectal cancer progression
Metabolic specialization structures gut bacterial niches and drives colorectal cancer progression
Xu, L.-L.; Seelbinder, B.; Zhou, Z.; Kuo, T.-H.; Sae-Ong, T.; Treibmann, S.; Damerell, V.; Brobeil, A.; Richter, K. M.; Mueller, M.; Toriola, A. T.; Shibata, D.; Li, C. I.; Byrd, D. A.; Figueiredo, J. C.; Hardikar, S.; Zielinski, C. E.; Bleckmann, A.; Ni, Y.; Correia-Melo, C.; Zimmermann, M.; Ulrich, C. M.; Gigic, B.; Panagiotou, G.
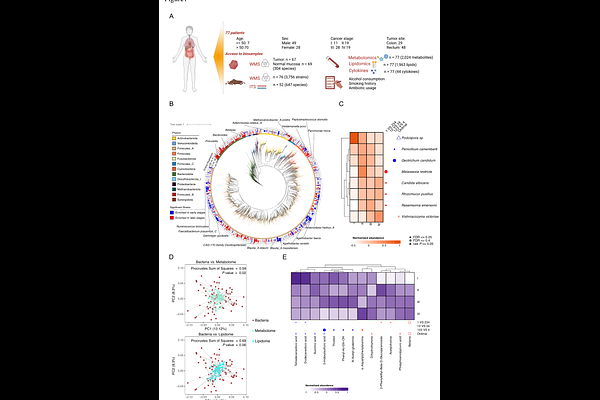
AbstractDespite the established association between the gut microbiome and colorectal cancer (CRC), the functional distinction between microbial passengers and drivers of CRC progression remains unresolved. Here, we collected stool, blood, as well as paired tumor, and normal mucosa tissues from seventy-seven CRC patients to characterize the systemic and localized impact of the gut microbiome on early- and late-stage CRC. By deep shotgun metagenomic sequencing, we identified distinct bacterial species and functions residing in tumor versus normal mucosa, highlighting an enrichment of oral-associated bacteria in tumor tissues. Several of these species remained undetected in the stool microbiome analysis. We further combined bacterial culturing with untargeted metabolomics of bacteria enriched in tumor and normal mucosa tissues, revealing distinct clusters of metabolic potential. Functional testing of multiple members from one cluster comprising both tumor- and mucosa-enriched species revealed Leptotrichia wadei as a pro-tumorigenic bacterium in a murine CRC model. Single-nucleus RNA sequencing and in vitro experiments further demonstrated that L. wadei and its secretome induces M2 macrophage polarization to promote tumor growth. Overall, our study shows that metabolic specialization structures microbial colonization niches, while species-specific metabolic outputs identify functional drivers of CRC progression, and uncovers L. wadei as an oncogenic bacterium in CRC.