Origin and evolution of the bread wheat D genome
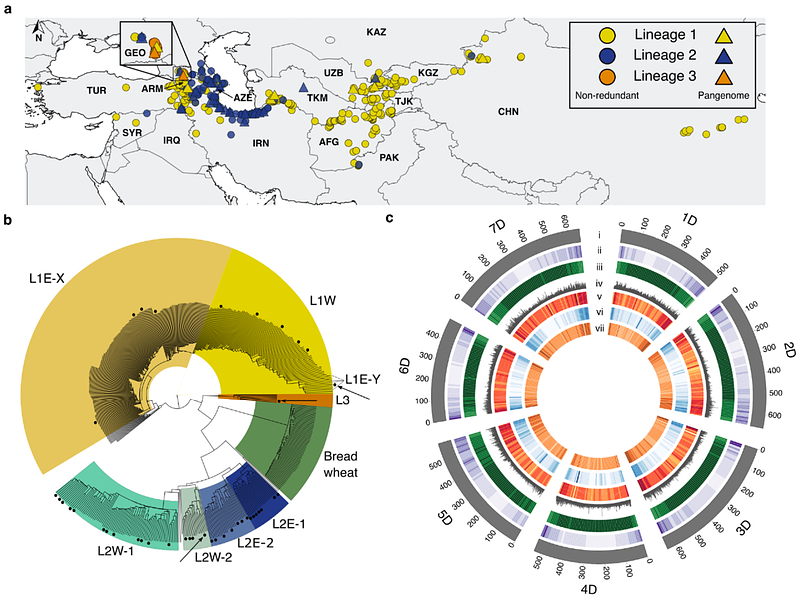
Origin and evolution of the bread wheat D genome
Cavalet-Giorsa, E.; Gonzalez-Munoz, A.; Athiyannan, N.; Holden, S.; Salhi, A.; Gardener, C.; Quiroz-Chavez, J.; Rustamova, S. M.; Elkot, A. F.; Patpour, M.; Rasheed, A.; Mao, L.; Lagudah, E. S.; Periyannan, S. K.; Sharon, A.; Himmelbach, A.; Reif, J. C.; Knauft, M.; Mascher, M.; Stein, N.; Chayut, N.; Ghosh, S.; Perovic, D.; Putra, A.; Perera, A. B.; Hu, C.-Y.; Yu, G.; Ahmed, H. I.; Laquai, K. D.; Rivera, L. F.; Chen, R.; Wang, Y.; Gao, X.; Liu, S.; Raupp, W. J.; Olson, E. L.; Lee, J.-Y.; Chhuneja, P.; Kaur, S.; Zhang, P.; Park, R. F.; Ding, Y.; Liu, D.-C.; Li, W.; Nasyrova, F. Y.; Dvorak, J.;
AbstractBread wheat (Triticum aestivum) is a globally dominant crop and major source of calories and proteins for the human diet. Compared to its wild ancestors, modern bread wheat shows lower genetic diversity caused by polyploidisation, domestication, and breeding bottlenecks. Wild wheat relatives represent genetic reservoirs, harbouring diversity and beneficial alleles that have not been incorporated into bread wheat. Here, we establish and analyse pangenome resources for Tausch\'s goatgrass, Aegilops tauschii, the donor of the bread wheat D genome. This new pangenome facilitated the cloning of a disease resistance gene and haplotype analysis across a complex disease resistance locus, allowing us to discern alleles from paralogous gene copies. We also reveal the complex genetic composition and history of the bread wheat D genome, involving previously unreported contributions from genetically and geographically discrete Ae. tauschii subpopulations. Together, our results reveal the complex history of the bread wheat D genome and demonstrate the potential of wild relatives in crop improvement.