Computational mapping of antibody-receptor energy landscapes to predict membrane internalization
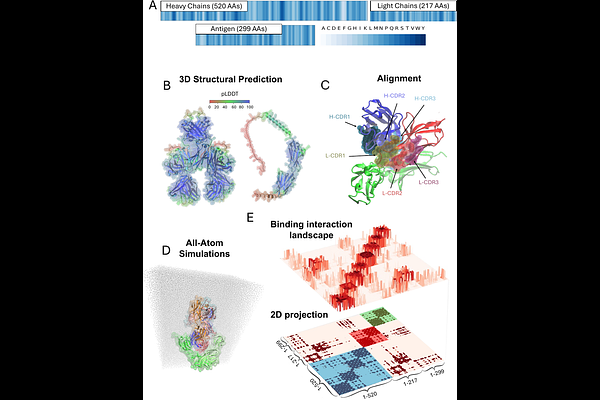
Computational mapping of antibody-receptor energy landscapes to predict membrane internalization
Llombart, P.; Nieto-Jimenez, C.; Pandiella, A.; Ocana, A.; Rene Espinosa, J.
AbstractAntibody internalization is a critical property for the development of antibody--drug conjugates and targeted therapeutics, yet antibody discovery pipelines typically prioritize binding affinity rather than functional internalization. Here we show that molecular dynamics simulations can map the binding energy landscape between antibody clones and the membrane-proximal receptor JAM-A, enabling computational predictions of tumor cell internalization. Using the sequences of newly generated anti--JAM-A monoclonal antibodies (mABs), we perform atomistic potential-of-mean force simulations to evaluate the binding free energy to the JAM-A receptor and their interaction fingerprint at residue-level resolution. We find that internalizing mAB clones exhibit a unique membrane-oriented contact topology that promotes cooperative receptor--receptor interactions, lowering the energetic barrier for early endocytic events. Reconstruction of the receptor binding energy landscape further reveals that electrostatic interactions between charged residues and multivalent cation-$\pi$ and polar interactions correlate with successful mAB internalization in ovarian cancer cells. In contrast, strong binding affinity of the fragment antigen-binding domain correlates with poor internalization. Together, our results establish molecular dynamics--guided clonal selection as a predictive framework for optimizing internalizing therapeutic antibodies and provide mechanistic insight into how antibody binding reshapes membrane-proximal receptor energetics to drive endocytosis.